Dr. Jia Yu
Research Focus
A perspective to understand the assembly of plant microbiota via metabolomic interactions between plant and bacteria
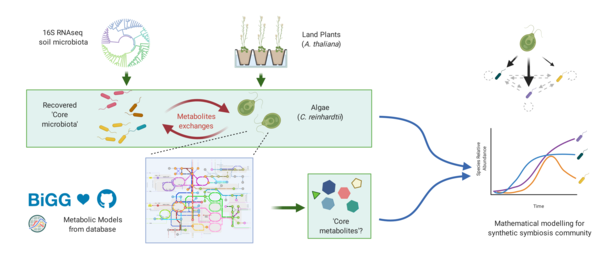
In nature, plants host a diverse community of microorganisms from different kingdoms. Among those microorganisms, bacteria were shown to play important roles in regulating the growth of plants (Durán et al., 2018). On the other side, the structure of the bacterial community can be shaped by plants and can be further stabilised at certain taxonomic level, furthermore, bacterial metabolites were found to potentially facilitate the community stabilisation (Trivedi et al., 2020, Goldford et al., 2018). Those results indicated that the behaviours of the bacterial community could also be shaped by plant cell metabolites and the community dynamics could also possibly be predicted (Qian and Akçay, 2020). To further investigate that hypothesis, establishing a synthetic method is required for wet-lab experiments (Fu et al., 2020). Algae can be a good substitute for plants by forming a similar microenvironment to host bacterial communities and they are suitable for high-throughput experiment designing (Seymour et al., 2017). We try to use Chlamydomonas reinhardtii as model organisms to study the bacteria growth with resources supplied from algae. It will help us to have a better understanding of: 1) how the bacterial community changes by knowing the metabolites consumption patterns for the bacteria 2) the possibility of choosing a highly simplified core microbiota but maintain the ecological properties.
Dr. Jia Yu

Department of Plant Microbe Interactions
Max Planck Institute for Plant Breeding Research