Dr. Concetta De Quattro
Research focus
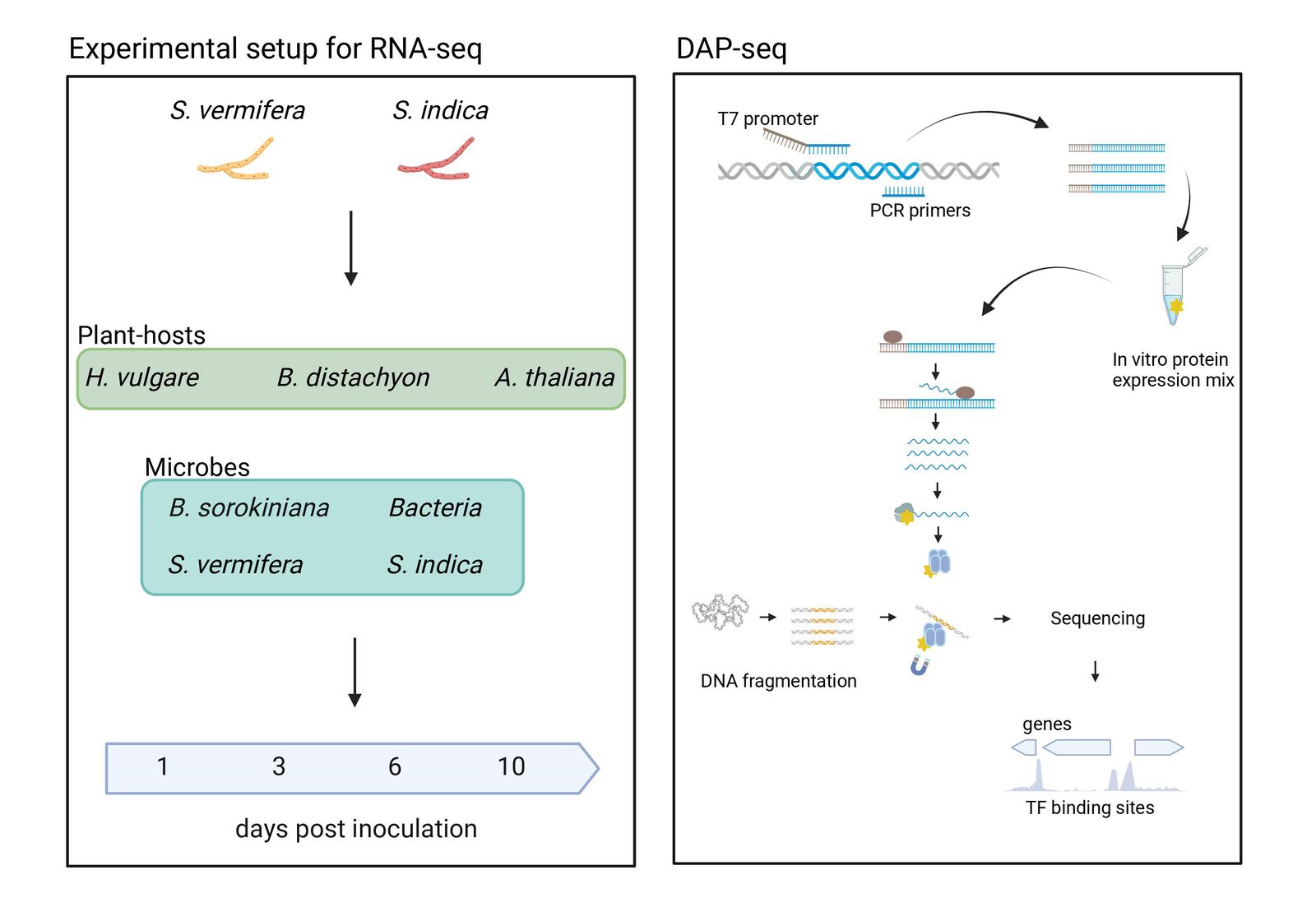
In the past, research has focused on identifying and clarifying the function of effectors of pathogenic fungi during colonization of a plant host species. While this has provided important insights into plant immunity and microbial infection strategies, it is now clear that these interactions between plants and microbes are inherently more complex and largely determined by multipartite interactions. This has led to the hypothesis that intermicrobial competition and cooperation are critical processes affecting plant-microbe interactions in the rhizosphere and may be controlled by microbial effector proteins. However, the molecular functions of effector proteins that specifically influence intermicrobial interactions are still largely unclear. Knowledge of how beneficial fungi regulate effector gene expression and function in different plant hosts and in response to other microbes is also lagging. Recent studies have revealed the role of various transcription factors, chromatin-based control, and effector epistasis in regulating effector expression in the host plant. Here, we propose to use DAP-seq and RNA-seq to generate cistrometes for Serendipita indica and S. vermifera, two closely related beneficial fungal symbionts from the order Sebacinales that inhabit the roots of phylogenetically diverse plant species to study the regulation of effector gene expression.