How to start a change in leaf ontogeny for a more efficient photosynthesis?
Photosynthesis is the process used by plants (and some other organisms like bacteria) to convert light energy into chemical energy, by producing sugars from carbon dioxide (CO2) and water (H2O). Photosynthesis is divided into two stages, the light reactions where light is harvested into the plant cells and the dark reactions where CO2 is fixed into sugars. The light reactions use H2O to produce the energy-rich molecules ATP and NADPH, which are fed as a substrate to the dark reactions.
Three major metabolic pathways for photosynthesis are known: C3, C4 and CAM. From these three modes of photosynthesis, C4 is the most efficient. This is because whenever the rate of the light reactions exceeds the rate of the dark reactions, the available CO2 within the leaf cannot keep up with the supply of ATP and NADPH from the light reactions. In that situation, the enzyme that fixes the CO2 by using the ATP and NADPH (Rubisco) will then use the extra ATP and NADPH to fix O2 starting the wasteful process of photorespiration. In C4-plants, there is barely any photorespiration, because they have a C4-cycle to concentrate CO2 near Rubisco so it never gets short of CO2.
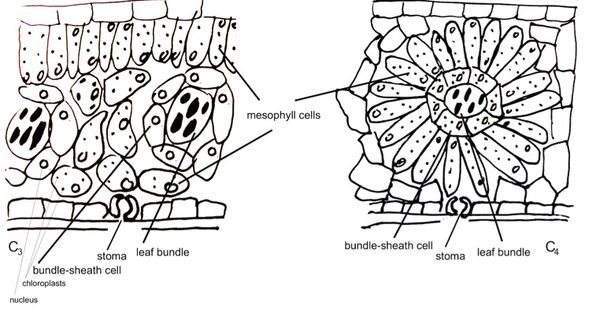
So why then are not all plants C4-plants? This is because C3-photosynthesis has arisen first during evolution and C4-photosynthesis has evolved out of C3-photosynthesis. Many researchers have been and are trying to understand the evolutionary pathway to evolve from a C3 into a C4-plant. A major contribution to the C4-cycle is the different leaf ontogeny in C4-plants, which is the sequence of developmental stages through which a leaf passes during growth. C4 plants develop highly specialized bundle-sheath cells around the leaf bundles (see figure). Changing the leaf ontogeny in a C3-plant towards C4 is seen as the first step to evolve C4-photosynthesis.
Within CEPLAS, we are trying to genetically induce this first step to evolve C4-photosynthesis. Because in history, C4-photosynthesis has evolved multiple times independently from C3-photosynthesis, it is thought that the necessary genes to activate the process are already present in C3-species. They just have to get activated in the right manner. In order to achieve this, we activate genes in the genome of a C3-species by inserting a bundle-specific promoter (isolated from a C4-species) randomly in the genome of the C3-species. We search for plants with aberrant leaf bundle ontogeny, scan the genome for the insertion location of the bundle-specific promoter, look which endogenous gene was affected by this insertion, and finally find out how it causes the aberrant bundle ontogeny.
Dr. Roxanne van Rooijen, Developmental and Molecular Plant Biology, Heinrich Heine University Düsseldorf
Planter’s Punch
Under the heading Planter’s Punch we present each month one special aspect of the CEPLAS research programme. All contributions are prepared by our young researchers.
Further Reading
Schuler ML, Mantegazza O, Weber AP, (2016) Engineering C4 photosynthesis into C3 chassis in the synthetic biology age. The Plant Journal 87, 51-65 [Abstract]
Engelmann S, Wiludda C, Burscheidt J, Gowik U, Schlue U, Koczor M, Streubel M, Cossu R, Bauwe P, Westhoff P (2008) The gene for the P-subunit of glycine decarboxylase from the C4 species Flaveria trinervia: analysis of transcriptional control in transgenic Flaveria bidentis (C4) and Arabidopsis (C3). Plant Physiology 146, 1773-1785 [Abstract]
About the Author

The article was written by Roxanne van Rooijen, who is a member of the CEPLAS Postdoc Programme. She aims to identify genes that control leaf anatomy and understand how they affect the photosynthetic performance of the leaf.