Communication between cells via peptides and receptors
Peptides and receptors in communication between plant cells
Plant cells communicate with each other using various signaling molecules such as phytohormones, sugars, and peptides. We mainly focus on small signaling peptides and their receptors. Once these signaling peptides are perceived by their receptors located at the plasma membrane (cell surface), signals are transduced into the cells and affect cellular activities, such as cell fate determination. In the last decades, a great number of studies have revealed that many secreted peptides are involved in numerous biological processes in plants such as stomatal patterning, stem cell maintenance, nutrients sensing, nodulation, and immune responses. In the Arabidopsis thaliana genome (widely used as a model plant), over 900 putative signaling peptides are predicted. However, most of them are uncharacterized peptides. Correspondingly, there are more than 600 predicted receptor like kinases in Arabidopsis genome and most of their functions are unknown as well.
What is an ovule?
The ovule is a part of a plants reproductive organ which contains the egg cell. After pollination, pollen tubes are attracted by secreted peptides and grow towards the egg cells in the ovules to enable fertilization. After fertilization, seeds develop as the products of fertilized and matured ovules. Therefore, the ovule is one of the most important organs regarding the plants reproductive success. At the same time, the number of ovules determines the maximum number of seeds. During the long domestication processes of crop plants, plant breeders have selected useful traits to improve crop productivity. Rice, barley and wheat are good examples. Nowadays, we get a lot of benefits from these improvements. For example, please consider your breakfast: Maybe you will realize that our daily life fully depends on seeds. Can you imagine your life without beer? (As you know well, beer is produced from barley grains or wheat in case of Weizenbier.) And of course, rice is crucial for Japanese people as their principle food as well as a main material for rice wine. From this aspect, the ovule is not only important for plants, but also important for us.
Cell-cell communication in ovule spacing
Within CEPLAS, we focus on the mechanism underlying ovule spacing using Arabidopsis thaliana as a model system. By genetic screening, we found genes which encode a secreted peptide and its receptor responsible for ovule density. These genes are found even in basal land plants, but are especially well conserved in Brassica species to which A. thaliana belongs. Until today, many genes were identified as responsible for ovule development, however the mechanisms which control ovule density and number of ovules are poorly understood. To address these questions, we combined molecular genetics (including genome editing), tissue clearing, confocal 3D imaging, and biochemical peptide-receptor interactions. Our analyses suggest that two pathways act in an antagonistic manner to control ovule density. One pathway generates more space between ovules and the other pathway reduces space between ovules. By controlling these pathways, we might be able to produce plants with higher ovule density, higher number of ovules and therefore higher number of seeds. We believe our studies in ovule spacing have potential ability to improve crop productivity in the future.
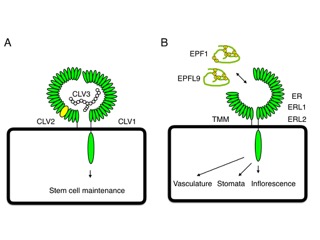
Figure 1: Signaling peptides are perceived by their receptors and transduce signals into cells. These specific peptide (ligand)-receptor interactions generate signals which determine cellular activities. Here are two examples shown which are well characterized. For details please see original articles. A: One is CLAVATA signaling pathway which control stem cell maintenance. CLAVATA1 (CLV1) and CLAVATA2 (CLV2) are a leucine rich repeat receptor kinase and receptor protein, respectively. CLAVATA3 (CLV3) is a signaling peptide and known as ligand of these receptors. B: The other is ERECTA (ER) signaling pathway which control various plant developmental processes including stomatal differentiation, inflorescence architecture, vascular development etc. ER and ER LIKE1 (ERL1) and ERL2 function redundantly and these receptors form a receptor complex with the receptor protein TOO MANY MOUTH (TMM). The cysteine rich peptides, EPIDERMAL PATTERNING FACTOR (EPF)/ EPIDERMAL PATTERNING FACTOR LIKE (EPFL) peptides are known as ligands of these receptors to control various processes.
Figure 2: Ovule (Seed) spacing in commercially available greens in super market. From top to bottom: snow peas, green beans, and bush beans. Each variety has different distances between its seeds (indicated by arrows). While snow peas have high seed density, the other two have low density. To increase productivity, the control of the distances between ovules is a key aspect to study.
Dr. Nozomi Kawamoto, Institute for Developmental Genetics, Heinrich Heine University Düsseldorf
Planter’s Punch
Under the heading Planter’s Punch we present each month one special aspect of the CEPLAS research programme. All contributions are prepared by our young researchers.
Further Reading
Clark, S.E., Williams, R.W., and Meyerowitz, E.M. The CLAVATA1 gene encodes a putative receptor kinase that controls shoot and floral meristem size in Arabidopsis. Cell, 1997, 89, 575-585 Abstract
Fletcher, J.C., Brand, U., Running, M.P., Simon, R., and Meyerowitz, E.M. Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems. Science, 1999, 283, 1911-1914 Abstract
Shpak, E.D., McAbee, J.M., Pillitteri, L.J., and Torii, K.U. Stomatal patterning and differentiation by synergistic interactions of receptor kinases. Science, 2005, 309, 290-293 Abstract
Lee, J.S., Kuroha, T., Hnilova, M., Khatayevich, D., Kanaoka, M.M., McAbee, J.M., Sarikaya, M., Tamerler, C., and Torii, K.U. Direct interaction of ligand-receptor pairs specifying stomatal patterning. Genes Dev., 2012, 26, 126-136 Abstract