Dr. St. Elmo Wilken
Research focus
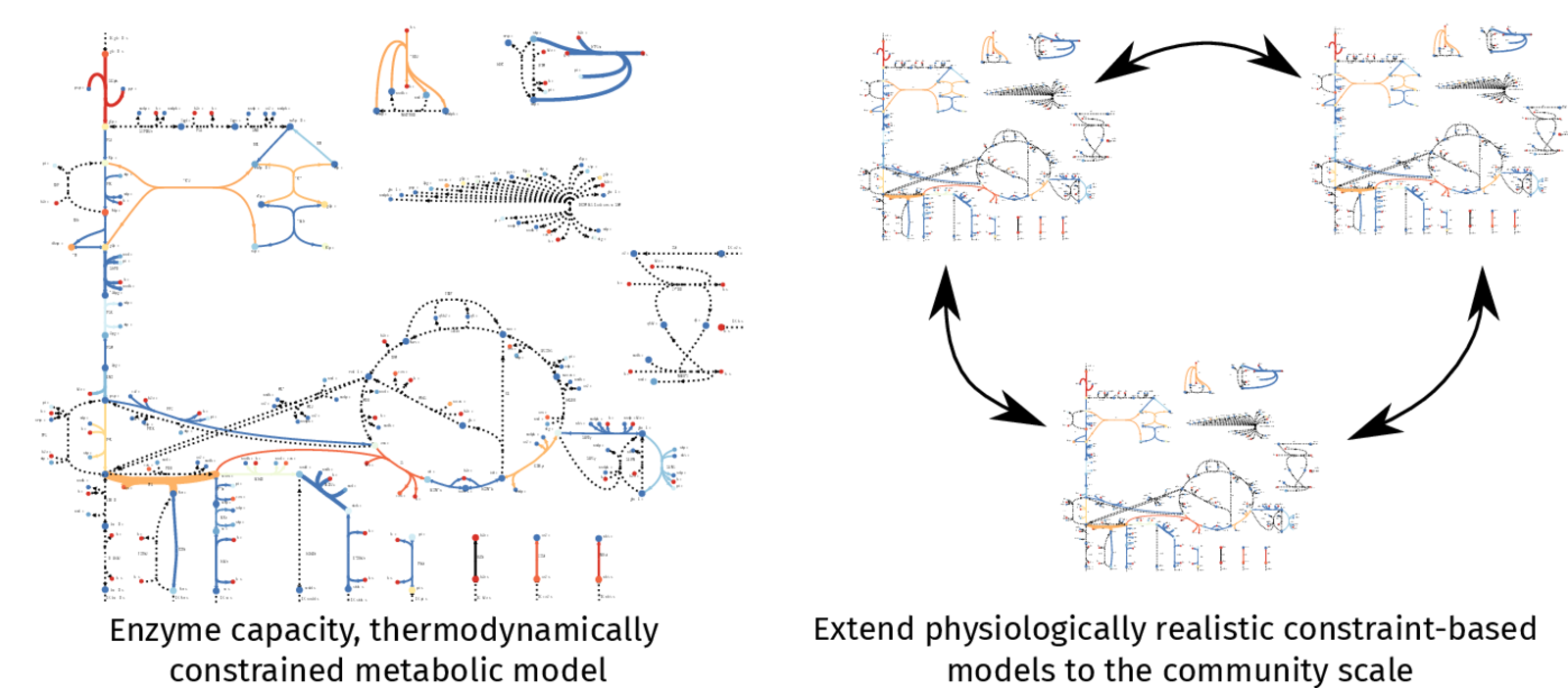
Constraint-based metabolic models are used to mechanistically describe the metabolism of a single organism at the genome-scale. By integrating omics data, phenotypically accurate predictions in diverse environments can be made to facilitate systematic strain engineering. Extending this framework to model microbiomes introduces more complexity, but could unlock insights into the general principles that shape microbial community composition and stability. In my project I will use metabolic models to investigate the design rules that affect microbial communities and apply engineering principles to tailor these communities for biotechnological purposes.
Dr. St. Elmo Wilken

Institute for Quantitative and Theoretical Biology
Heinrich Heine University Düsseldorf